-Search query
-Search result
Showing 1 - 50 of 52 items for (author: lou & zy)

EMDB-34314: 
SARS-CoV-2 RNA E-RTC complex with RMP-nsp9 and GMPPNP
Method: single particle / : Yan LM, Huang YC, Ge J, Liu ZY, Gao Y, Rao ZH, Lou ZY

EMDB-34316: 
SARS-CoV-2 E-RTC bound with MMP-nsp9 and GMPPNP
Method: single particle / : Yan LM, Rao ZH, Lou ZY

EMDB-34310: 
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Method: single particle / : Yan LM, Rao ZH, Lou ZY

EMDB-34311: 
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Method: single particle / : Yan LY, Huang YC, Rao ZH, Lou ZY

EMDB-34312: 
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Method: single particle / : Yan LM, Huang YC, Ge J, Liu ZY, Gao Y, Rao ZH, Lou ZY

EMDB-34313: 
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Method: single particle / : Yan LM, Huang YC, Ge J, Liu ZY, Gao Y, Rao ZH, Lou ZY
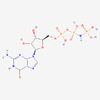
EMDB-34317: 
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Method: single particle / : Yan LM, Huang YC, Ge J, Liu ZY, Gao Y, Rao ZH, Lou ZY

EMDB-34308: 
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Method: single particle / : Yan LM, Rao ZH, Lou ZY
EMDB-34318: 
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Method: single particle / : Yan LM, Huang YC, Ge J, Liu ZY, Gao Y, Rao ZH, Lou ZY

EMDB-33126: 
Cryo-EM structure of Dot1L and H2BK34ub-H3K79Nle nucleosome 1:1 complex
Method: single particle / : Ai HS, Liu AJ, Lou ZY, Liu L

EMDB-33127: 
Cryo-EM structure of Dot1L and H2BK34ub-H3K79Nle nucleosome 2:1 complex
Method: single particle / : Ai HS, Liu AJ, Lou ZY, Liu L
EMDB-33128: 
cryo-EM map of Dot1L and H2BK34ub-H3K79Nle nucleosome complex containing addition map (1:1)
Method: single particle / : Ai HS, Liu AJ, Lou ZY, Liu L
EMDB-33131: 
cryo-EM structure of H2BK34ub nucleosome
Method: single particle / : Ai HS, Liu AJ, Lou ZY, Liu L

EMDB-33132: 
cryo-EM structure of unmodified nucleosome
Method: single particle / : Ai HS, Liu AJ, Lou ZY, Liu L
EMDB-33139: 
cryo-EM map of Dot1L and H3K79Nle nucleosome complex (active state)
Method: single particle / : Ai HS, Liu AJ, Lou ZY, Liu L
EMDB-33141: 
cryo-EM map of Dot1L and H3K79Nle nucleosome complex (inactive state)
Method: single particle / : Ai HS, Liu AJ, Lou ZY, Liu L

EMDB-23312: 
BG505 SOSIP.v5.2 in complex with VRC40.01 and RM19R Fabs
Method: single particle / : Cottrell CA, Torres JL, Wu NR, Ward AB

PDB-7lg6: 
BG505 SOSIP.v5.2 in complex with VRC40.01 and RM19R Fabs
Method: single particle / : Cottrell CA, Torres JL, Wu NR, Ward AB

EMDB-31138: 
Co-transcriptional capping machineries in SARS-CoV-2 RTC: Coupling of N7-methyltransferase and 3'-5' exoribonuclease with polymerase reveals mechanisms for capping and proofreading
Method: single particle / : Yan LM, Yang YX, Li MY, Zhang Y, Zheng LT, Ge J, Huang YC, Liu ZY, Wang T, Gao S, Zhang R, Huang YY, Guddat LW, Gao Y, Rao ZH, Lou ZY

EMDB-23424: 
Cryo-EM map of Q23.17_DS-SOSIP in complex with Glycan276-Dependent Broadly Neutralizing Antibody 179NC75 Fab
Method: single particle / : Manne K, Acharya P

PDB-7llk: 
Cryo-EM structure of Q23.17_DS-SOSIP in complex with Glycan276-Dependent Broadly Neutralizing Antibody 179NC75 Fab
Method: single particle / : Manne K, Acharya P

PDB-7l6o: 
Cryo-EM structure of HIV-1 Env CH848.3.D0949.10.17chim.6R.DS.SOSIP.664
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-30232: 
Structure of Dot1L-H2BK34ub Nucleosome Complex
Method: single particle / : Lou ZY, Liu L, Cao L, Ai HS, Sun ZX

EMDB-23518: 
Cryo-EM map of DH851.3 bound to HIV-1 CH505 Env
Method: single particle / : Edwards RJ, Manne K, Acharya P

PDB-7lu9: 
Cryo-EM structure of DH851.3 bound to HIV-1 CH505 Env
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23411: 
Cryo-EM map of BG505 DS-SOSIP in complex with glycan276-dependent broadly neutralizing antibody VRC40.01 Fab
Method: single particle / : Manne K, Acharya P

EMDB-23412: 
Cryo-EM map of BG505 DS-SOSIP in complex with Glycan276-Dependent Broadly Neutralizing Antibody VRC33.01 Fab
Method: single particle / : Manne K, Acharya P

PDB-7ll1: 
Cryo-EM structure of BG505 DS-SOSIP in complex with glycan276-dependent broadly neutralizing antibody VRC40.01 Fab
Method: single particle / : Manne K, Acharya P

PDB-7ll2: 
Cryo-EM structure of BG505 DS-SOSIP in complex with Glycan276-Dependent Broadly Neutralizing Antibody VRC33.01 Fab
Method: single particle / : Manne K, Acharya P

EMDB-23519: 
Cryo-EM map of DH898.1 Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer
Method: single particle / : Edwards RJ, Manne K, Acharya P

PDB-7lua: 
Cryo-EM structure of DH898.1 Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23152: 
Cryo-electron microscopy reconstruction of antibody DH898.1 Fab-dimer bound to glycans 332, 392, and 396 of HIV Env CH848 10.17 SOSIP trimer
Method: single particle / : Edwards RJ, Acharya P

EMDB-23153: 
Cryo-electron microscopy local refinement of antibody DH898.1 Fab-dimer bound to glycans 332, 392, and 396 of HIV Env CH848 10.17 SOSIP trimer
Method: single particle / : Edwards RJ, Acharya P

EMDB-23124: 
Cryo-electron microcospy reconstruction of CH848.3.D0949.10.17chim.6R.DS.SOSIP.664 HIV Env
Method: single particle / : Edwards RJ, Acharya P

EMDB-23145: 
Cryo-electron microscopy reconstruction of locally refined antibody DH898.1 Fab-dimer
Method: single particle / : Edwards RJ, Acharya P

PDB-7l6m: 
Cryo-EM structure of DH898.1 Fab-dimer from local refinement of the Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23094: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to one copy of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R, Acharya P

EMDB-23095: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to two copies of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R, Acharya P

EMDB-23097: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound domain-swapped antibody 2G12 from masked 3D refinement
Method: single particle / : Manne K, Henderson R, Acharya P

PDB-7l02: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to one copy of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R, Acharya P

PDB-7l06: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to two copies of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R, Acharya P

PDB-7l09: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound domain-swapped antibody 2G12 from masked 3D refinement
Method: single particle / : Manne K, Henderson R, Acharya P

EMDB-9607: 
Anthrax Toxin Receptor 1-bound spent particles of Seneca Valley Virus in acidic conditions
Method: single particle / : Lou ZY, Cao L

EMDB-3982: 
Molecular Architecture of the Jumonji C histone demethylase KDM5B.
Method: single particle / : Aduri NA, Gajhede M

EMDB-9608: 
Anthrax Toxin Receptor 1-bound full particles of Seneca Valley Virus in acidic conditions
Method: single particle / : Lou ZY, Cao L

EMDB-9611: 
Anthrax Toxin Receptor 1-bound the Seneca Valley Virus in neutral conditions
Method: single particle / : Lou ZY, Cao L

EMDB-9612: 
Structure of Seneca Valley Virus in acidic conditions
Method: single particle / : Lou ZY, Cao L

EMDB-9613: 
Structure of Seneca Valley Virus in neutral condition
Method: single particle / : Lou ZY, Cao L
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
